DNA and the Genetic Code
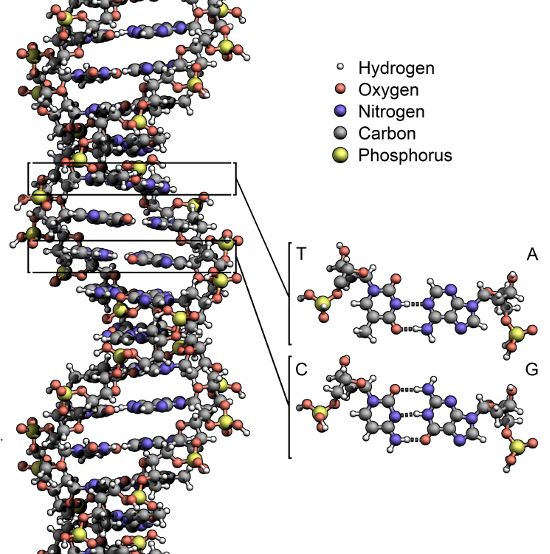
DNA resembles a twisted ladder. Each rung is a pair of nucleobases: adenine (A) always pairs with thymine (T), and cytosine (C) always pairs with guanine (G).
After a series of discoveries in the first half of the 20th century, scientists had figured out that chromosomes contain long molecules of deoxyribonucleic acid (DNA), and that genes are stored within these molecules. Then, in 1953, a team of two scientists—American geneticist James Watson and British biophysicist Francis Crick—successfully determined the molecular structure of DNA, a feat for which they later earned a Nobel Prize. It turned out that DNA is shaped like a double helix. It resembles a twisted ladder, and genetic information is encoded in the “rungs” of the ladder. Each rung is called a base pair and consists of two of the following four chemical compounds, the names of which are abbreviated with the letters A, T, C, and G: adenine (A), thymine (T), cytosine (C), and guanine (G). These four compounds, called nucleobases (or simply bases, for short), are the alphabet in which genetic information is written. We can think of a gene as a sequence of the letters A, T, C, and G, analogous to the sequences of ones and zeros used in computer programming.
With a few exceptions, each cell in a living organism’s body contains a copy of the same set of DNA molecules.Exceptions include red blood cells, which lose their DNA as they mature; also gametes, which contain only half the genome, as previously noted. That set of DNA molecules is called the organism’s genome. For example, the human genome consists of 92 DNA molecules, paired together in 46 chromosomes, which in turn are grouped in 23 pairs. Our genome stores a mind-boggling amount of information. Altogether, the DNA molecules in a single human cell contain over 3 billion base pairs. If you unraveled all the DNA from one cell of your body and stretched it out, it would be about 2 meters long!
The information in DNA is encoded using a programming language known as the genetic code. The “words” and symbols of this programming language are called codons. Each codon consists of three nucleobases arranged contiguously along one side of a DNA molecule. For example, the sequence “ATG” (spelled using the bases adenine, thymine, and guanine, in that specific order) is a start codon, indicating the beginning of a new sentence in the programming language. Similarly, the letters “TGA” form a stop codon, which functions as the period at the end of the sentence. Still other codons function as words representing the various molecular building blocks used to construct proteins and other molecules needed by the cell, as discussed below.
Each gene or allele of Mendel’s theory corresponds to a sentence in the language of the genetic code, written on one or moreSome genes consist of a single, contiguous segment of DNA. However, many genes are located in multiple non-contiguous segments. See the fine-print section, below, for further explanation. segments of DNA. Genes have many functions, some of which are not yet well-understood, but the most common genes contain information used to assemble proteins—large molecules that perform specialized functions or serve as the custom-made components of various structures in the cell. Proteins are built from up to 21 different kinds of simple molecules called amino acids, 20 of which are common to all living organisms. The genetic code represents each amino acid with one or more codons. For example, the codons CAA and CAG both represent an amino acid called glutamine, which has chemical formula C5H10N2O3.
So, to recap, the genetic code is a programming language in which the alphabet consists of nucleobases and the “words” are three-letter codons. Each gene is a “sentence” in that programming language, spelling out instructions for the assembly and operation of the molecular components within a living cell. The mechanisms involved in carrying out these instructions are truly marvelous, but I won’t go into all the details here. For just a taste of the magnificent complexity and efficiency of these processes, see the video below, which explains how molecular machines copy genetic code from DNA onto a molecule called messenger RNA (mRNA), trim off superfluous segments, then transmit the edited information via mRNA to ribosomes—molecular factories which act as miniature 3D printers, following the instructions to assemble specific types of proteins. Nearly all of the complicated machinery in a living cell is made of specialized protein molecules constructed by ribosomes. Even ribosomes themselves are made of numerous protein parts, which were constructed by other ribosomes!
This three-minute video explains and illustrates the intricate mechanisms involved in carrying out the instructions encoded in DNA.
With the help of molecules called transfer RNA (tRNA), a ribosome assembles a protein by joining a sequence of amino acids together in the precise order specified by the codons in a gene. This long chain of amino acids then folds into a protein of a specific 3-dimensional shape. (Proteins typically fold themselves due to the electromagnetic forces and chemical interactions between neighboring amino acids, though some are assisted in the folding process by other mechanisms in the cell.) Thus, the shape and functional characteristics of the protein are determined by the amino acid sequence specified by a gene—that is, by a corresponding sequence of codons on a strand of DNA.
A crucial role of the genetic information in DNA is to specify all the diverse structures of proteins, which serve as the mechanical parts of microscopic engines, robots, and other amazing nanotechnology employed within the cell. But where did all this genetic information come from in the first place? The origin of the very first genes and gene processing mechanisms is disputed, as we’ll see later in this chapter and in the following chapter. However, there is at least one natural process by which some new genes might be created: mutation. We’ll consider that on the next page.
This interactive simulation allows you to experiment with gene expression—the complex process by which information stored in DNA is used to produce proteins. Open the “Expression” experiment and look for the “Biomolecule Toolbox” near the upper left corner of the window. It contains several biomolecules and molecular machines you can experiment with: transcription factors (molecular switches that turn genes on or off), RNA polymerase (a machine that transcribes digital information from DNA to create a strand of mRNA), ribosomes (molecular factories that follow the instructions in mRNA to assemble proteins), and mRNA destroyers (you can guess what those do). The simulation also includes a strand of DNA with three genes, which you can access by clicking the “Next Gene” and “Previous Gene” buttons to scroll back and forth along the DNA. Try attaching various transcription factors to the regulatory region at the start of each gene, along with RNA polymerase, and see what happens with various combinations. If you succeed in producing mRNA strands, try dragging ribosomes into the simulation to create proteins. To open the simulation in a new tab, click here.